Bird fingers confuse me, but the explanations confuse me more, it seems.
I didn’t mean to post today, but I’ve just read a new review/hypothesis paper about the identities of the stunted little things that pass for fingers in the wings of modern birds. The review part is fine, but I’m not sure I get the difference between the hypothesis Čapek et al. (2013) are proposing and the hypothesis they are trying to replace/improve.
To recap: the basic problem with bird fingers is that fossil, genetic and developmental evidence seem to say different things about them.
1. Fossils: birds pretty clearly come from dinosaurs, and the early dinosaurs we have fossils of have five fingers on their hands with the last two being reduced. Somewhat closer to birds, you get four fingers with #4 vestigial. And the most bird-like theropods have only three fingers, which look most like digits 1, 2 and 3 of your ordinary archosaur. (Although Limusaurus messes with this scheme a bit.)
2. Embryology: in developing limb buds, digits start out as little condensations of tissue, which develop into bits of cartilage and then finger bones. Wing buds develop a short-lived condensation in front of the first digit that actually forms, and another one behind the last “surviving” digit. Taking this at face value, then, the fingers are equivalent to digits 2, 3 and 4.
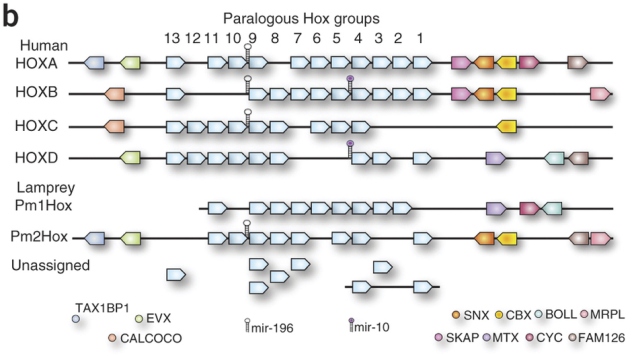
3. Genetics: In five-fingered limbs, each digit has a characteristic identity in terms of the genes expressed during its formation. The first finger of birds is most like an ordinary thumb, both when you focus on individual genes like members of the HoxD cluster and when you take the entire transcriptome. However, the other two digits have ambiguous transcriptomic identities. That is, bird wings have digit 1 and two weirdos.
Add to this the fact that in other cases of digit loss, number one is normally the first to go and number four stubbornly sticks around to the end, and you can see the headache birds have caused.
So those are the basic facts. The “old” hypothesis that causes the first part of my confusion is called the frame shift hypothesis, which suggests that the ancestors of birds did indeed lose digit 1, as in the digit that came from condensation 1 – but the next three digits adopted the identities of 1-2-3 rather than 2-3-4. (This idea, IMO, can easily leave room for mixed identities – just make it a partial frame shift.)
Čapek et al.’s new one, which they call the thumbs down hypothesis, is supposedly different from this. This is how the paper states the difference:
The FSH postulates an evolutionary event in which a dissociation occurs between the developmental formation of repeated elements (digits) and their subsequent individualization.
versus
According to the TDH no change of identity of a homeotic nature occurs, but only the phenotypic realization of the developmental process is altered due to redirected growth induced by altered tissue topology. Digit identity stays the same. Also the TDH assumes that the patterning of the limb bud, by which the digit primordia are laid down, and their developmental realization, are different developmental modules in the first place.
(Before this, they spent quite a lot of words explaining how the loss of the original thumb could trigger developmental changes that make digit 2 more thumb-like.)
I…. struggle to see the difference. If you’ve (1) moved a structure to a different position, (2) subjected it to the influence of different genes, (3) and turned its morphology into that of another structure, how exactly is that not a change in identity?
Maybe you could say that “an evolutionary event” dissociating digit formation and identity is different from formation and identity being kind of independent from the start, but I checked Wagner and Gauthier’s (1999) original frame shift paper, and I think what they propose is closer to the second idea than the first:
Building on Tabin’s (43) insight, we suggest causal independence between the morphogenetic processes that create successive condensations in the limb bud and the ensuing developmental individualization of those repeated elements as they become the functional fingers in the mature hand, thus permitting an opportunity for some degree of independent evolutionary change.
Am I missing something? I feel a little bit stupid now.
***
References:
Čapek D et al. (2013) Thumbs down: a molecular-morphogenetic approach to avian digit homology. Journal of Experimental Zoology B, published online 29/10/2013, doi: 10.1002/jez.b.22545
Wagner GP and Gauthier JA (1999) 1,2,3 = 2,3,4: A solution to the problem of the homology of the digits in the avian hand. PNAS 96:5111-5116