It’s kind of hard to begin this post. First of all, let’s get the important news out of the way: I’ve just published a paper. In a moment, I’ll get around to discussing it at even more than my usual length, but I feel that I can’t do my excited puppy act without at least trying to capture how bloody much this paper means to me. The following may get a little personal; if you want to jump straight to the Cool Stuff, feel free to scroll a couple of paragraphs down.
<personal bit>
As you may have guessed from the long silence here, it’s not been a good handful of years, Real Life and mental health-wise. After my PhD, the prospect of the research career I’d dreamed of since I first began to grasp the meaning of the word “scientist” no longer seemed so dreamlike. It may surprise you to hear this from someone who finished a PhD with four published papers and spent the years of said PhD blathering regularly on the internet, but I find writing things for other people to read very, very stressful. In the case of a job application or a thesis chapter, that becomes “I’m not eating or sleeping properly” stressful. (Don’t ask me how I survived 20+ years of formal education.)
Long story short, for the last 3 years I’ve been getting by with a minimum wage job for which I’m both vastly overqualified and singularly ill-suited. I started the research project that culminated in the paper you can now read (for free, yay!) in BMC Evolutionary Biology (Szabó and Ferrier, 2018) while unemployed and broke, and I did most of it in my free time around work. This paper is a hard-won victory over myself and my circumstances. It’s a tiny glint of self-worth in the depth of the tunnel. In some ways, it was harder than my thesis: no funding body to satisfy, no lab mates to gripe at, no deadlines to spur me on. The only constant was my ex-supervisor turned co-author, who took my hobby project under his wings for the slim reward of having his name on a paper and nudged me into finishing it with unending patience. Here’s to Dave Ferrier, champion of non-model organisms, homeobox guy extraordinaire and all-round excellent human being. Dave, I hope you know you’re an absolute star.
</personal bit>
With that out of the way, it’s time for the Cool Stuff. There are Hox genes! More Hox genes than anyone ever imagined! (That is kind of the point, in fact!)
Apologies for the word count. I thought it would be a good idea to explain a few things, but also, I think I enjoy waffling about my baby far too much 😊
Hox therapy
The story of my Hox paper begins with an unemployed biologist with an overabundance of free time and a desperate need to do something scientific. Since I have a slightly odd idea of “fun”, back in 2015 I decided to catalogue Hox gene (or rather, protein) diversity in the animal kingdom, with particular focus on obscure and poorly studied groups. (I didn’t get very far, as we’ll see.)
Since it’s hard to discuss the paper without dropping some arcane zoological nomenclature, here’s my trusty old animal phylogeny to (re)acquaint us with the general outlines of the animal kingdom (I might need to update this in light of the Great Ctenophore Controversy some day, but we’re not dealing with anything outside the Bilateria today):

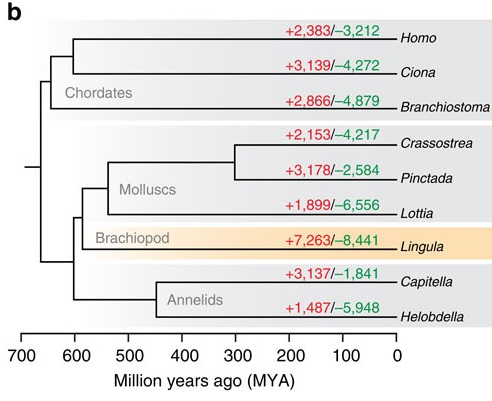
For the purposes of my paper, we’re zooming into the deuterostome branch, which looks something like this on the inside (borrowing my own rather lacklustre last-minute figure from Szabó and Ferrier [2018]):

Everything on this tree apart from chordates (that’s us) belongs to a group called Ambulacraria, which contains two phyla, hemichordates (top two branches) and echinoderms (the next five). Echinoderms are the more familiar of the two – starfish and sea urchins and suchlike – and also the focus of my project. (I could find no Hox gene data from pterobranchs, which puts a slight caveat on everything I say about hemichordates)
Back to Hox genes.
Hox genes were kind of my gateway drug into evolutionary developmental biology. A few decades earlier, they had served the same purpose for developmental biology as a whole, since they were among the first genes to be discovered that (1) directed embryonic development (2) were comparable between very disparate animal groups. The short version, which will suffice for our purposes here, is that Hox genes are important in what we eggheads call anteroposterior patterning, or determining what body parts go where along the head (anterior) to tail (posterior) axis of a (bilaterian) animal.
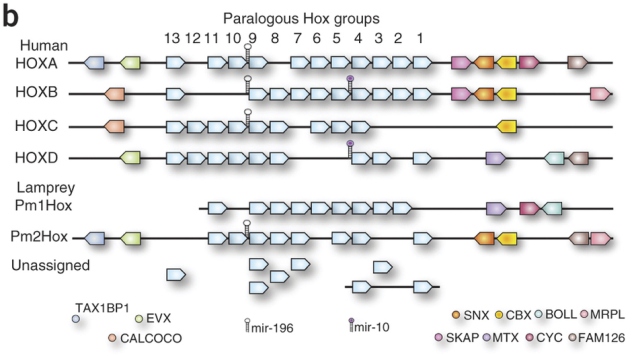
In (I think, I haven’t counted) the majority of animals that have them, Hox genes are clustered to a greater or lesser extent. Rather than being scattered haphazardly across the genome, they sit close to one another along the same stretch of DNA. (Duboule [2007] is an excellent – albeit now slightly out of date – review of the various known configurations.)
Since my study is about echinoderms, the schematic Hox cluster shown below is the neatest known example from an echinoderm, the crown-of-thorns starfish Acanthaster planci (source: Baughman et al., 2014):

In this image, Hox genes are colour-coded according to a commonly used classification scheme. This classification is mostly based on the homeodomain, or the “business” end of the protein that a Hox gene encodes. A homeodomain makes up a relatively small portion (maybe 1/5th on average) of a typical Hox protein, but it’s the part that interacts with the DNA switches through which Hoxes control their target genes, and it’s often the only part that is similar enough to be compared between different Hox types.
The important genes for us today are the “posterior” Hox genes shown in pink and red above, especially the last two. The four posterior Hox genes seen here represent the “standard” set for ambulacrarians, although it’s uncertain whether Hox11/13b-c were already separate genes or just a single precursor gene in the ambulacrarian ancestor.
Eureka… or WTF?
“The most exciting phrase to hear in science, the one that heralds new discoveries, is not Eureka! (I found it!) but rather, ‘hmm… that’s funny…” – Almost certainly not Isaac Asimov
In creating my grand catalogue, I’d quickly breezed through vertebrates (which are all essentially the same for my purposes) and other chordates (for which the data I could find were rather limited). I thought echinoderms would be an easy job, too: there were good in-depth studies of a few species, and they hadn’t revealed anything terribly unusual other than a rearrangement of the Hox cluster in sea urchins (Cameron et al., 2006).
In fact, through comparison with their sister group, the hemichordates (Freeman et al., 2012), it seemed likely that the ancestral echinoderm had a nice, ordered Hox cluster with few if any oddities (Baughman et al., 2014). So I clicked my way to the wonderful Echinobase, which has searchable draft genomes from four of the five living classes of echinoderms (crinoids, a.k.a sea lilies and feather stars, are missing, although a genome in a very early, fragmentary stage exists here). I expected to double-check the published data, collect the same genes from the groups for which Hox papers hadn’t been published, and be off to protostomes in a day or two. Two years later, I still haven’t made it to protostomes, but I’ve gone rather deeper than expected in echinoderms…
(Below: my cast. The main characters are Strongylocentrotus purpuratus [photo: Kirt L. Onthank] and Lytechinus variegatus [photo: Hans Hillewaert] representing sea urchins, Patiria miniata [photo: Jerry Kirkhart] and Acanthaster planci [photo: JSLUCAS75] for sea stars, Parastichopus parvimensis [from here] and Apostichopus japonicus [photo: OpenCage] for sea cucumbers, Metacrinus rotundus [photo: OpenCage] and Anneissia japonica [photo: OpenCage] for crinoids, Ophiothrix spiculata [photo: Jerry Kirkhart] for brittle stars, with supporting acts from Peronella japonica [sea urchins, photo: Endo et al., 2018], Ophiopsila aranea [brittle stars, photo: Bernard Picton], Balanoglossus simodensis [photo: Misaki Marine Biological Station, U of Tokyo], Saccoglossus kowalevskii [photo: Lowe lab] and Ptychodera flava [photo: Moorea BioCode via CalPhotos] for hemichordates, and Branchiostoma floridae [photo via JGI genome portal], Latimeria menadoensis [photo: Claudio Martino] and Callorhinchus milii [photo: fir0002/Flagstaffotos] for chordates. I sourced the photos through Wikipedia/Wikimedia Commons where I could; other sources are linked where applicable.)

You see, I didn’t want to stop at just homeodomains. Homeodomains are cool and important and all, but one thing I’d learned from my earlier forays into the world of Hox genes was that valuable information hid in small patches of conserved sequence elsewhere in their proteins. Besides, I am a pathological perfectionist. I felt a terrible need to collect complete Hox sequences wherever possible.
I already mentioned that sequence similarity between Hoxes outside the homeodomain can be weak to non-existent. I ran into this problem with Echinobase’s brittle star, Ophiothrix spiculata. Using the known sea urchin Hoxes to search its genome, I’d found believable matches for many of them, but the 11/13s defeated me. I had two homeodomains that I thought represented 11/13b and c, but I couldn’t for the life of me recover the rest of the proteins.
The problem with genome databases (or their great advantage depending on your perspective) is that they contain all of the DNA that could be sequenced from the owner of the genome. The problem with Hox genes – most of our genes, in fact – is that they aren’t continuous stretches of DNA. Your typical gene exists in multiple segments (exons) separated by a whole lot of DNA that leaves no trace in the protein product of the gene. (Hox genes normally have two or three exons, the first of which is devoid of homeodomain parts.)
When a gene is expressed, the cell first makes an RNA copy of all that, which is edited to throw out the introns and splice the exons together. That intron-less RNA copy is then carried off to be translated into a protein. Transcriptomes are derived from the RNA copies of active genes. Introns lie forgotten on the cutting room floor: in the sequenced transcripts, one exon continues straight into the next. Therefore, if I could find a brittle star transcriptome, and the 11/13b-c homeodomains in it, perhaps there would be enough of the rest in there to reconstruct those elusive first exons.
Luckily, Delroisse et al. (2016) had published exactly what I needed. In one of their transcriptomes, I found a homeodomain that looked like my Ophiothrix Hox11/13c, as part of a near-complete sequence. Excited, I did the reciprocal search against the Ophiothrix genome…
… and hit neither 11/13b nor 11/13c.
So here I am, staring at a beautiful match between this transcript and a part of the Ophiothrix genome that I hadn’t examined before. The match contains sequence from the first exon, which, given my previous experience with these buggers, is a sure sign that they’re the same gene. And it’s neither of the ones I’d expected.
A bit later in a different database, I hit upon an automatically predicted sea urchin protein that definitely isn’t 11/13b or c either. This is the model sea urchin, S. purpuratus, the one I thought we knew inside out when it came to Hoxes. I check the genome on Echinobase, and lo and behold, there’s the third 11/13b-c type gene, and it’s nowhere near the Hox cluster.
If memory serves, it’s roughly at this point that the words, “What. The. Actual. Fuck. Is. Going. On.” occur in my research notes. (Complete with punctuation.)
I checked the other species on Echinobase. Three 11/13b-c genes again, every time. Over on Genbank, I found a complete protein sequence from a sand dollar that Tsuchimoto and Yamaguchi (2014) had previously classified as 11/13c by exclusion. The Japanese duo had a clear b, but this other sequence was behaving oddly in their phylogenetic analyses. Now I had the obvious explanation: it wasn’t 11/13c at all.*
I wrote to Dave and found out that this was also news to him. By all appearances, I had stumbled on something truly new, in a gene family that’s both iconic in our field, and dear to my obsessive little heart.
We decided to try to turn it into a paper.
In search of the alphabet’s end
Once we’d made that decision, and following Dave’s advice, I had a few tasks ahead of me. I had to check how far back in evolution our new gene (which we called Hox11/13d) went. I had to test whether it had truly escaped the Hox cluster in all of our study species. I had to refresh my memory on deuterostome posterior Hox genes in general, both for paper-writing purposes and in case there was a forgotten reference to our “new” gene lurking somewhere in the literature.
There wasn’t, but.
In a figure legend in Thomas-Chollier et al., 2010), there is a brief mention of an unnamed “Hox11/13c-like” sequence in sea urchins. When I saw that, I damn near soiled myself, but the authors couldn’t definitively identify this sequence as a Hox gene, so they left it at that throwaway comment and a few bits of supplementary data. Luckily, they had a gene ID that I could look up on Echinobase.
Gods help me, it turned out to be another new Hox. When the shock of Hox11/13d had barely worn off, I was confronted with a possible Hox11/13e. And this one wasn’t in the Hox cluster either.
Aside from not being part of the Hox cluster, Hox11/13d is a pretty good echinoderm Hox gene. The homeodomain it encodes is reminiscent of Hox11/13b and c, and, although they are hard for automated searches to find, there are similarities outside the homeodomain that place it firmly in the same group as b-c.
Unlike d, Thomas-Chollier’s “11/13c-like” sequence isn’t that 11/13c-like at all, as you might have guessed from the fact that they weren’t even sure it’s a Hox. The region immediately following the homeodomain (sometimes known as the C-peptide) is very similar to the same part of Hox11/13d. These kinds of motifs can sometimes be used to tell different Hox genes apart. Two C-peptides being strongly similar is a clue that we’re dealing with related genes. However, the homeodomain of Hox11/13e, as we indeed dubbed Thomas-Chollier’s sequence, is really, really weird. It isn’t just unlike 11/13c, it’s unlike anything else I’d seen before. It groups with posterior Hoxes when we test it against a variety of homeodomains, but you wouldn’t know that simply from looking at it.
It is, however, an oddball with a history. As strange as that homeodomain is, once I knew what I was looking for, I found examples in all my other echinoderms. This combination of strong conservation of one Hox gene with considerable differences from other Hox genes just screams “study me more!”, especially when you realise that Hox11/13e appears to be limited to echinoderms (unless something like it is hiding in protostomes…). I looked quite carefully in the hemichordates available to me (Simakov et al., 2015), but the only thing I found that wasn’t one of the “canonical” four posteriors is something called “Abdominal B-like”, which is weird in its own way and not obviously connected to either of our two new genes.
Tangled histories and unhelpful clues
I alluded to the question of Hox11/13b-c origins earlier on. Posterior Hox genes in deuterostomes are notoriously difficult to classify (Ferrier et al., 2000; Thomas-Chollier et al., 2010). When you try to use traditional tree-building methods on them, you get a big unresolved mess, as if the twigs on the tree emerged from an impenetrable mist that hides the arrangement of the older branches from view. Ambulacrarians are definitely the better-behaved half of the Deuterostomia in this regard, since we can say with some confidence that Hox9/10, 11/13a and at least a single precursor to 11/13b-c were present in their last common ancestor.
Nonetheless, two new genes, at least one of which is clearly close to 11/13b-c, complicate matters (Abdominal B-like, as they say in scientist-speak, is beyond the scope of this work). Were they lost in hemichordates? Did echinoderms undergo extra gene duplications, and if so, was it from one or two ancestral genes? Where on earth does Hox11/13e fit? I did a lot of exploratory tree-building for this paper, none of which was particularly helpful in answering those questions.
My other hope was to look at the parts of the protein sequence that led me to my new Hoxes in the first place: all the stuff other than the homeodomain. Using a program called MEME, I found a fair few conserved motifs, but they only seemed to add to the confusion. Hox11/13e, for which I only had first exons (and tentative ones at that) from sea urchins and sea stars, yielded nothing of use apart from its striking C-peptide. In the others, the distribution of motifs created a patchwork of similarities that didn’t neatly align with any one possible history. Echinoderm Hox11/13c mostly did its own thing, while b and d each shared a different subset of motifs with one or both of the hemichordate b-c proteins.
I’m almost inclined to think that there was a single, “prototype” Hox11/13b+ sequence in the ambulacrarian ancestor, which contained all of the motifs I found. In that scenario, separate b and c (and d and maybe e) genes would have evolved independently in hemichordates and echinoderms, and each descendant gene would have lost some of the original motifs more or less at random. Duplicated genes can split the functions of their single ancestor between them (Force et al., 1999), so why not motifs? Short sequence motifs like the ones I was looking for can have important functions, after all. It’s a possibility, but we may never know for sure.
Hox genes gone rogue
I mentioned before that Hox11/13d was outside the Hox cluster. Well, so is Hox11/13e. As far as I can tell, Hox 11/13d and e always reside on separate chunks of the genome form any other Hox gene, including each other. They are always accompanied by neighbouring genes that aren’t Hoxes. Although detachment of a posterior gene from an otherwise apparently intact Hox cluster also happened in ragworms (Hui et al., 2012), it’s still a surprise in echinoderms. Since the relationship between the organisation of Hox genes and their regulation in space and time is… kinda complicated, we can’t really tell what, if anything, all this wandering implies without actually looking at some gene expression.
What are they for?
Then there’s the question of what on earth these genes do. Thanks to Tsuchimoto and Yamaguchi (2014), we know that Hox11/13d is active in later embryonic stages of some sea urchins. It even looks like it might be working with Hox11/13b in a Hox-like fashion, the two of them having adjacent expression domains. We have some transcriptomic evidence that this gene is also active in other sea urchins, brittle stars and starfish, but no idea what it’s doing in any of the above.
We know even less about Hox11/13e. The only evidence for expression I’m aware of is from starfish testicles, and testicles will express any old piece of DNA with an “on” switch. If it’s somehow involved in development, it must be either at very low levels that are difficult to capture in a transcriptome, or at developmental stages that weren’t included in the data I encountered.
If it does have a role in adult echinoderm development, that would be crazy exciting, as both adult echinoderm anatomy and Hox11/13e are so weird and unique. Although they develop from bilaterally symmetrical larvae, adult echinoderms have dispensed with the symmetry that gave Bilateria its name. Instead, like a sea anemone (or a regular anemone…), they are radially symmetrical. Hox genes are involved in both larval and adult development in echinoderms, but from what little I’ve been able to glean from the existing literature, it’s different subsets in larvae and adults rather than the entire Hox cluster together. Is Hox11/13e in the “adult” subset, missed until now due to its unusual sequence? I really hope someone with a lab and a ready supply of baby echinoderms investigates in the near future…
A lesson about expectations
I could go on for a lot longer about this project, but it’s probably time to form some sort of conclusion. For me, perhaps the most important take-home message of this adventure is not what I found, but how and where and why I found it.
I didn’t set out to discover anything. All I wanted to do was collect and organise information already out there. (If a genie popped out of my desk lamp, I might just wish for a full-time job where I get to build my Hox directory… given the volume of genome data already out there and coming out every time I look, continuing this as a hobby project in my free time seems hopelessly Sisyphean now.)
The discovery of Hox11/13d and all that followed was an accidental side effect of my penchant for perfectionism. If I’d contented myself with the homeodomains most students of Hox evolution focus on, I would never have seen a Hox that wasn’t in the books, a Hox I hadn’t expected to exist.
Expectations are important. I’d told myself that I wanted to make sure I had everything, but when my searches spat out a hundred different results, I started to slack off soon after I ticked off the Hoxes I knew. I gave the rest of the hit list a half-hearted effort at best. Hox11/13d has a homeodomain that’s split across two exons, and Hox11/13e is weird. In a search that scores both the closeness and the length of a match, that pushes them to the bottom of the results, where a casual observer, or an observer who thinks they know what they’re looking for, will most likely miss them. I thought I knew that sea urchins had a single, intact(ish) Hox cluster with 11 genes. I’d read a pretty good paper on it. Only the paper wasn’t quite right, after all.
To me, this study stands as a reminder to keep looking. In an era when new genomes are popping up left and right and Big Data with automated analyses is the scientific zeitgeist, it’s still worth rolling your sleeves up, picking up the old magnifying glass and taking a closer look – even in organisms you think you know. You might just chance upon some real treasure.
***
Note:
*A “Hox11/13c” behaving oddly should be immediately suspicious based on what I saw in my own trees, where echinoderm Hox11/13c consistently formed a strongly supported group. But that’s hindsight for you…
***
References:
Baughman KW et al. (2014) Genomic organization of Hox and ParaHox clusters in the echinoderm, Acanthaster planci. Genesis 52:952-958
Cameron RA et al. (2006) Unusual gene order and organization of the sea urchin hox cluster. JEZ B 306:45-58
Delroisse J et al. (2016) De novo adult transcriptomes of two European brittle stars: spotlight on opsin-based photoreception. PLoS ONE 11: e0152988
Duboule D (2007) The rise and fall of Hox gene clusters. Development 134:2549-2560
Endo M et al. (2018) Hidden genetic history of the Japanese sand dollar Peronella (Echinoidea: Laganidae) revealed by nuclear intron sequences. Gene 659:37-43
Ferrier DEK et al. (2000) The amphioxus Hox cluster: deuterostome posterior flexibility and Hox14. Evol Dev 2:284-293
Force A et al. (1999) Preservation of duplicate genes by complementary, degenerative mutations. Genetics 151:1531-1545
Freeman R et al. (2012) Identical genomic organization of two hemichordate Hox clusters. Curr Biol 22:2053-2058
Hui JH et al. (2012) Extensive chordate and annelid macrosyntheny reveals ancestral homeobox gene organization. Mol Biol Evol 29:157-165
Simakov O et al. (2015) Hemichordate genomes and deuterostome origins. Nature 527:459-465
Szabó R and Ferrier DEKF (2018) Two more Posterior Hox genes and Hox cluster dispersal in echinoderms. BMC Evol Biol 18:203
Thomas-Chollier M et al. (2010) A non-tree-based comprehensive study of metazoan Hox and ParaHox genes prompts new insights into their origin and evolution. BMC Evol Biol 10:73
Tsuchimoto J and Yamaguchi M (2014) Hox expression in the direct-type developing sea urchin Peronella japonica. Dev Dyn 243:1020-1029